Plant substances as potential medicines
Purpose:
The purpose of this lab is to gather native plants and create extracts in order to determine their potential antimicrobial abilities.
Materials:
- balance, weight boat, lab scoops
- LB broth base
- media bottles, 250 mL
- sterilizer/autoclave
- water bath, 37 degrees Celsius
- sterile LB agar
- laminar flow hood and disinfectant
- plastic safety glasses
- Bunsen burner and gas lighter
- inoculating loop, Ni/Cr wire
- petri dishes, 60 x 15 mm, sterile
- E. coli JM109 (stock plate)
- plant specimen
- mortar and pestle
- pipet, 10 mL and pump
- short-stemmed plastic funnels
- filter paper disks, 5 mm diameter
- Beakers, 100 mL
- syringe, 10 mL and filter, 0.2 microliters
- reaction tubes and rack, 1.7 mL
- absolute methanol
- pipet, 1 mL and pump
- dry block heater/heat block
- fine-tipped forceps
- ampicillin
- glass spreader
- incubator oven, 37 degrees Celsius
- forceps
- alcohol
- clamp
Procedure:
Gather plants and crush them into a paste with water. Pour the paste into a filter and filter out the excess plant matter until all you're left with is plant extract in a tube. Repeat with methanol. Pipette each extract into a sterile microcentrifuge tube and leave the methanol tube open to evaporate excess methanol. Place small filter papers in the centrifuge tube and allow them to soak in the extract. Take a petri dish filled with agar and draw separating lines for two MeOH quadrants and two water quadrants, and label a + and - in the center for control filters. Pipette E coli onto your plate and spread it with a flame-sterilized streaking tool. Place your filters in the corresponding areas using flame-sterilized tweezers. Wait 24 hours and observe the resulting bacterial growth.
Results:
The purpose of this lab is to gather native plants and create extracts in order to determine their potential antimicrobial abilities.
Materials:
- balance, weight boat, lab scoops
- LB broth base
- media bottles, 250 mL
- sterilizer/autoclave
- water bath, 37 degrees Celsius
- sterile LB agar
- laminar flow hood and disinfectant
- plastic safety glasses
- Bunsen burner and gas lighter
- inoculating loop, Ni/Cr wire
- petri dishes, 60 x 15 mm, sterile
- E. coli JM109 (stock plate)
- plant specimen
- mortar and pestle
- pipet, 10 mL and pump
- short-stemmed plastic funnels
- filter paper disks, 5 mm diameter
- Beakers, 100 mL
- syringe, 10 mL and filter, 0.2 microliters
- reaction tubes and rack, 1.7 mL
- absolute methanol
- pipet, 1 mL and pump
- dry block heater/heat block
- fine-tipped forceps
- ampicillin
- glass spreader
- incubator oven, 37 degrees Celsius
- forceps
- alcohol
- clamp
Procedure:
Gather plants and crush them into a paste with water. Pour the paste into a filter and filter out the excess plant matter until all you're left with is plant extract in a tube. Repeat with methanol. Pipette each extract into a sterile microcentrifuge tube and leave the methanol tube open to evaporate excess methanol. Place small filter papers in the centrifuge tube and allow them to soak in the extract. Take a petri dish filled with agar and draw separating lines for two MeOH quadrants and two water quadrants, and label a + and - in the center for control filters. Pipette E coli onto your plate and spread it with a flame-sterilized streaking tool. Place your filters in the corresponding areas using flame-sterilized tweezers. Wait 24 hours and observe the resulting bacterial growth.
Results:
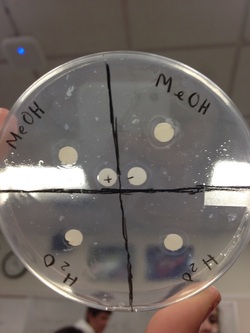
All extracts on my plate were unsuccessful in inhibiting microbial growth. The MeOH extracts had clear rings around them but also had bacterial growth right next to the filters anyways, meaning they were ineffective. The rings around the MeOH samples were about 2mm thick.
The water samples were also ineffective, though the rings around them were about 1.5mm thick.
Both controls worked successfully; water had bacteria right up next to it and the Ampicillin filter had no bacterial growth around it.
If I were to do this experiment again, I might remove the excess liquid on the filters by touching it to the side of the microcentrifuge tube or onto another sterile filter, where capillary action will remove excess liquid. This is to prevent the excess liquid from clearing away the streaked bacteria and creating confusing results, as you can see with the rings in my experiment. I would even repeat my experiment with exactly the same factors otherwise, considering my partner got two positives while I got two negative results.
The water samples were also ineffective, though the rings around them were about 1.5mm thick.
Both controls worked successfully; water had bacteria right up next to it and the Ampicillin filter had no bacterial growth around it.
If I were to do this experiment again, I might remove the excess liquid on the filters by touching it to the side of the microcentrifuge tube or onto another sterile filter, where capillary action will remove excess liquid. This is to prevent the excess liquid from clearing away the streaked bacteria and creating confusing results, as you can see with the rings in my experiment. I would even repeat my experiment with exactly the same factors otherwise, considering my partner got two positives while I got two negative results.
Analysis:
Neither of my extracts returned a positive, while my partner, who had the same samples as I, got one positive and one weak-positive. This could mean that I made a mistake in applying the filters or in spreading the bacteria (unlikely), or perhaps that her results returned a false-positive. The bacteria could have been cleared away initially by the extra fluid on the filter papers, not allowing the bacteria to colonize. This would explain the rings around my negative samples, too.
TLAB Questions:
If an extract gives a negative result in the antimicrobial assay, this does not mean that the extract is not an antimicrobial agent. The extract/filters could be improperly prepared, compromising the experiment, or the could only kill certain types of bacteria. The bacteria could even exhibit immunity to the agent. If some of the methanol extracts smell like alcohol, this means that the methanol has not fully evaporated. As alcohol is already a known antimicrobial substance, this can compromise the results of your experiment. In order to identify exactly which compound exhibits antimicrobial action in an extract, you can separate and test individually the compounds through chromatography, which separates individual molecules based on polarity.
Neither of my extracts returned a positive, while my partner, who had the same samples as I, got one positive and one weak-positive. This could mean that I made a mistake in applying the filters or in spreading the bacteria (unlikely), or perhaps that her results returned a false-positive. The bacteria could have been cleared away initially by the extra fluid on the filter papers, not allowing the bacteria to colonize. This would explain the rings around my negative samples, too.
TLAB Questions:
If an extract gives a negative result in the antimicrobial assay, this does not mean that the extract is not an antimicrobial agent. The extract/filters could be improperly prepared, compromising the experiment, or the could only kill certain types of bacteria. The bacteria could even exhibit immunity to the agent. If some of the methanol extracts smell like alcohol, this means that the methanol has not fully evaporated. As alcohol is already a known antimicrobial substance, this can compromise the results of your experiment. In order to identify exactly which compound exhibits antimicrobial action in an extract, you can separate and test individually the compounds through chromatography, which separates individual molecules based on polarity.
